Entry Version:
Citation:
Pancreapedia: Exocrine Pancreas Knowledge Base, DOI: 10.3998/panc.2021.15
| Attachment | Size |
|---|---|
| 355.14 KB |
1. General Functioning of the mTOR Pathway
The mTOR signaling pathway is a nutrient sensing mechanism coupled to mTOR, the mammalian or mechanistic target of rapamycin (23, 36, 52). mTOR is the target of the anti-fungal metabolite rapamycin, which is named after the island Rapa Nui (Easter Island) from whose soil it was first isolated and which has broad antiproliferative and immunosuppresive properties (51). Genetic screens in the early 1990’s in yeast identified two genes TOR1 and TOR2 that mediated the effects of rapamycin. Biochemical studies then led to the identification of the mammalian form (12, 43). mTOR is an atypical protein kinase related to phosphoinositide 3- kinase family although it is a Ser/Thr targeted kinase and not a lipid kinase. It is a large protein of about 2,500 amino acids with multiple domains including a C terminal kinase domain and a FKBP-rapamycin binding (FRB) domain (4).
mTOR is a component of two complexes, TORC1 and TORC2; each contains other proteins, some of which are shared. However, TORC1 uniquely contains the scaffolding protein Raptor (regulatory associated protein of mammalian target of rapamycin) (27) and PRAS40 (proline rich Akt substrate of 40 kDa) (59), whereas TORC2 contains, among other components, Rictor (rapamycin-insensitive companion of mTOR) (36). Both Raptor and PRAS40 are inhibitory proteins; phosphorylation blocks this inhibition. PRAS40 represents an essential component for insulin activation of TORC1 (61). Raptor is an essential component and its genetic deletion leads to loss of TORC1 activity (7).
Much less is known as to the functions of TORC2. It is a regulator of the actin cytoskeleton in both yeast and mammalian cells (65). More recent studies have shown it to be active in phosphorylating various protein kinases especially Akt (24).
Only TORC1 is acutely sensitive to rapamycin which inhibits some, but not all, of TORC1 functions (41). This inhibition requires FKBP-12 (FK506 binding protein of 12 kDa). Much is known about the function of TORC1 which mediates the growth promoting effects including protein synthesis, lipid synthesis, inhibition of autophagy, ribosome and lysosome biogenesis, and energy metabolism (61). The effects on protein synthesis are mediated in part by activation of S6K1 (S6 Kinase-1) which phosphorylates ribosomal protein S6 and in part, by activation of initiation and elongation translation factors, the latter through elongation factor 2 kinase (41). In addition, TORC1 phosphorylates 4E-BP1, a binding protein which, upon phosphorylation, releases the initiation factor eIF4E which acts as a mRNA cap binding protein (60). The overall rate of protein synthesis also depends on the number of ribosomes, and TORC1 enhances the synthesis of ribosomal proteins and RNA (41). TORC1 promotes lipid synthesis by activating SREBP (sterol responsive element binding protein) (51).
The action of TORC1 to inhibit autophagy is mediated by phosphorylation of ULK1 (unc-51 like autophagy activating Kinase-1) and Atg13 (autophagy related 13) which blocks the formation of the phagosome (39, 53). Recently, TORC1 has also been shown to regulate the abundance of proteasomes; when TORC1 is inhibited, the ability of proteasomes to degrade proteins increases (1, 42, 55).
A variety of upstream signals regulate TORC1 including growth factors (and some hormones that stimulate growth of their target cells), amino acids, oxygen levels, and glucose (Figure 1). Growth factors and insulin reflect the fed state of the organism and promote anabolic processes. Binding of insulin to its receptor activates PI3K and Akt that are upstream of TORC1, Akt phosphorylates and inhibits TSC1/2 the tuberosclerosus complex which acts as a tumor suppressor by acting as a GAP for Rheb (Ras homolog enriched in brain), a GTPase which is one of the major activators of TORC1. Akt also phosphorylates PRAS40 and thereby relieves its inhibition of mTOR in TORC1. TORC1 limits its own activation by the negative feedback of S6K1 on the early steps in PI3K activation especially by insulin.
Figure 1. General and simplified diagram of mTORC1 pathway set in pancreatic acinar cell, where the pathway is regulated by CCK and insulin. Red arrows indicate activation; Black arrows indicate inhibition (which may involve multiple steps); Green arrow indicates translocation. Biological processes regulated by mTORC1 are shown at the bottom of the figure, along with the key proteins mediating the effect.
Another major signal regulating TORC1 activity is the abundance of amino acids which tells the organism to undergo anabolic activity (5, 31). Conversely, the absence of adequate amino acids is a stress which leads to the shutting down of biosynthetic pathways and the induction of autophagy. The major amino acids sensed are the branched chain amino acids, especially leucine, as well as arginine and glutamine. Amino acid sensing involves recruitment of TORC1 from cytoplasm to lysosomes where it interacts with proteins including the Rag GTPases (32), a protein complex termed the ragulator (44), amino acid transporters (26) and the lysosomal vacuolar ATPase (20). In the presence of amino acids, Rheb which is also localized on the lysosome, is activated by deubiquitination, and in turn, allosterically activates TORC1 kinase activity (14, 19). This mTORC1 activation by Rheb is inhibited by TSC, that converts Rheb from the GTP- to the GDP-bound state (2, 61). Amino acids are necessary for almost all other mechanisms activating TORC1 (3). Low oxygen or low glucose levels prevent TORC1 signaling through AMP Kinase and reduce the activity of the proteins REDD and BMIP3 (53, 63). DNA damage also inhibits TORC1. As a result of this network of interactions, growth and other anabolic activities can only take place in the presence of a supporting milieu. For more details see recent reviews (15, 33, 53).
2. mTOR Signaling in Pancreatic Cells
mTOR signaling in pancreas was first recognized and is most commonly followed through phosphorylation of the pathway’s downstream mediators, ribosomal protein S6 and 4E-BP1 (56). Phospho-specific antibodies are widely available for S6 (Figure 2A) and its upstream kinase, S6K1 which is activated by TORC1. 4E-BP1 resolves into multiple bands on Western blots with the higher band being most highly phosphorylated (Figure 2B) (48). Such measurements showed that in isolated rodent pancreatic acini CCK and similarly acting secretagogues (bombesin, carbachol) activated S6K1 (9), increased phosphorylation of S6, 4E-BP1 (9, 10, 50, 56), and EF2K (elongation factor 1 Kinase (50) and that these effects were blocked by rapamycin, the TORC1 inhibitor. Moreover, rapamycin blocked the increase in protein synthesis stimulated by CCK in isolated acini (10). These studies showed that the primary cell type involved in the exocrine pancreas is the acinar cell and this has been reinforced in vivo where CCK injection increased phosphorylation of S6 and 4E-BP1 as well as the phosphorylation of eIF4E and the formation of the eIF4F initiation complex (11). Elevating endogenous CCK by feeding the trypsin inhibitor camostat (17, 28) or diverting bile pancreatic juice (28) βamino acids activate the TORC1 pathway and leucine and other branched chain amino acids activate protein synthesis and the TORC1 pathway both in pancreas in vivo and in isolated pancreatic acini (29, 49, 58). Additionally, mTORC1 activity has been shown to be reduced after a short- (46), and long-term (16) feeding of a protein-free diet; demonstrating the involvement of dietary amino acids on its physiological stimulation.
Ribosomal protein S6 and 4E-BP1 signaling in the pancreas are also affected by insulin and diabetes (40, 45, 57). TORC1 signaling is not required for secretion of digestive enzymes (9) but is required for protein synthesis and adaptive growth (10, 17, 18, 46).
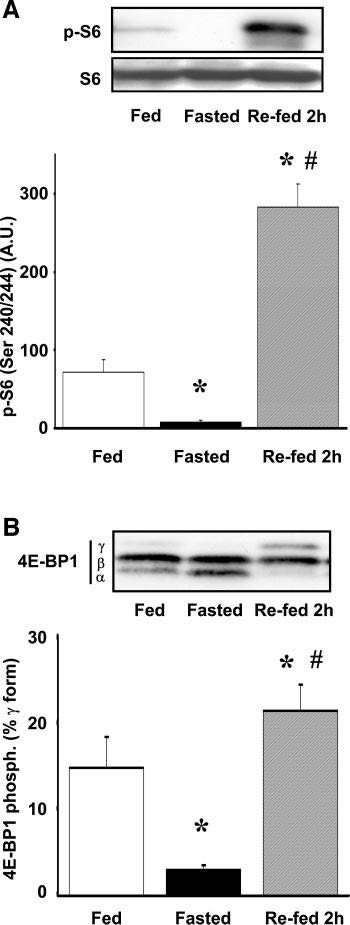
Figure 2. Effect of fasting and feeding on the phosphorylation of S6 (A) and eIF4E binding protein 4E-BP1 (B). In A, S6 phosphorylation is expressed as arbitrary units (A.U.), and each column is the means ± SE of 9–14 animals/group. Representative blots for S6 phosphorylation and total S6 are shown in the insets. In B, the phosphorylation levels of 4E-BP1 are represented as the amount of 4E-BP1 in the γ-phosphorylated form as a percentage of the total 4E-BP1. Inset: representative immunoblot with positions of α-, β-, and γ-forms of 4E-BP1 is noted on left. The most highly phosphorylated γ-form exhibits the slowest electrophoretic mobility and does not bind eIF4E. (From Ref. 48).
In addition to studies of acinar cells, the TORC1 pathway appears to play a role in activated pancreatic stellate cells where it mediates effects of insulin to enhance collagen synthesis and fibrosis (62). These in vitro effects of insulin were blocked by TOR inhibitors rapamycin and KU63794. TORC1 also plays a role in the endocrine pancreas where it is involved in islet development, beta cell growth and insulin processing and secretion (6, 8, 22, 37, 54).
The importance of the TORC1 pathway in the exocrine pancreatic response to feeding is shown by the activation of the downstream components when mice fasted overnight are refed (48). In this study, protein synthesis was also increased with feeding without a change in mRNA levels for digestive enzymes, indicating the importance of translational control primarily by the TORC1 pathway in synthesis of new digestive enzymes after secretion (48).
Similar effects have also been seen in neonatal pigs (25). TORC1 was also shown to play a role in the hypertrophic response to feeding a high protein diet and this was independent of CCK (18). Conversely, pancreatic atrophy was seen in response to a loss of TORC1 signaling when mice were fed a protein-free diet (16). These in vivo responses involve multiple hormones including CCK and insulin and nutrients acting directly, particularly amino acids (16, 46).
3. mTOR Signaling and Pancreatic Disease
mTOR signaling has been shown to be reduced during experimental pancreatitis, involved in the reduction of pancreatic digestive enzymes synthesis during the development of the disease (47). It has also been implicated in other disease states with altered growth and metabolism including cancer and diabetes as well as aging where the life extending effect of low-calorie diets is believed mediated by reduced TORC1 signaling (65). TORC1 activity is increased in many pancreatic ductal adenocarcinomas (PDAC) in part due to mutations in upstream regulatory molecules including PTEN, AKT and TSC1/2. Most PDAC cancers have RAS mutations leading to activation of the MEK/ERK pathway which can inactivate TSC1/2 and thereby activate TORC1 (13, 64). Rapamycin analogs have been considered as potential therapeutic agents for pancreatic cancer and other cancers. However, these inhibitors have not shown significant effects in single agent clinical trials, though individual patients have shown responses (30). Currently attention has focused on dual agent therapy as well as identifying patients with specific patterns of gene activation that may be more responsive. In this context, genetically modified mice with Ras mutation and PTEN deficiency show sensitivity to TORC1 inhibition in contrast to those with Ras and p53 mutations which are not sensitive (13, 38). Another study using a mouse model of decreased TSC1 by haploid sufficiency showed enhanced mTOR signaling and tumorigenesis could be blocked with dual inhibition of mTOR and MEK (13, 35).
When TSC1 was completely ablated in the embryonic pancreas, mice developed pancreatic acinar adenocarcinoma superimposed on atrophy of the normal pancreas (21, 34). This is a rare form of pancreatic carcinoma in humans with the tumor cells having acinar characteristics and expressing amylase. It is generally slower growing and less malignant than pancreatic ductal adenocarcinoma.
4. References
- Adegoke OAJ, Beatty BE, Kimball SR, and Wing SS. Interactions of the super complexes: When mTORC1 meets the proteasome. Int J Biochem Cell Biol 117: 105638, 2019. PMID: 31678320.
- Angarola B, and Ferguson SM. Coordination of Rheb lysosomal membrane interactions with mTORC1 activation. F1000Res 9: 2020. PMID: 32518628.
- Avruch J, Long X, Ortiz-Vega S, Rapley J, Papageorgiou A, and Dai N. Amino acid regulation of TOR complex 1. Am J Physiol Endocrinol Metab 296: E592-602, 2009. PMID: 18765678.
- Aylett CH, Sauer E, Imseng S, Boehringer D, Hall MN, Ban N, and Maier T. Architecture of human mTOR complex 1. Science 351: 48-52, 2016. PMID: 26678875.
- Bar-Peled L, and Sabatini DM. Regulation of mTORC1 by amino acids. Trends Cell Biol 24: 400-406, 2014. PMID: 24698685.
- Bartolome A, and Guillen C. Role of the mammalian target of rapamycin (mTOR) complexes in pancreatic beta-cell mass regulation. Vitam Horm 95: 425-469, 2014. PMID: 24559928.
- Bentzinger CF, Romanino K, Cloetta D, Lin S, Mascarenhas JB, Oliveri F, Xia J, Casanova E, Costa CF, Brink M, Zorzato F, Hall MN, and Ruegg MA. Skeletal muscle-specific ablation of raptor, but not of rictor, causes metabolic changes and results in muscle dystrophy. Cell Metab 8: 411-424, 2008. PMID: 19046572.
- Blandino-Rosano M, Barbaresso R, Jimenez-Palomares M, Bozadjieva N, Werneck-de-Castro JP, Hatanaka M, Mirmira RG, Sonenberg N, Liu M, Ruegg MA, Hall MN, and Bernal-Mizrachi E. Loss of mTORC1 signalling impairs beta-cell homeostasis and insulin processing. Nat Commun 8: 16014, 2017. PMID: 28699639.
- Bragado MJ, Groblewski GE, and Williams JA. p70s6k is activated by CCK in rat pancreatic acini. Am J Physiol 273: C101-109, 1997. PMID: 9252447.
- Bragado MJ, Groblewski GE, and Williams JA. Regulation of protein synthesis by cholecystokinin in rat pancreatic acini involves PHAS-I and the p70 S6 kinase pathway. Gastroenterology 115: 733-742, 1998. PMID: 9721171.
- Bragado MJ, Tashiro M, and Williams JA. Regulation of the initiation of pancreatic digestive enzyme protein synthesis by cholecystokinin in rat pancreas in vivo. Gastroenterology 119: 1731-1739, 2000. PMID: 11113094.
- Brown EJ, Albers MW, Shin TB, Ichikawa K, Keith CT, Lane WS, and Schreiber SL. A mammalian protein targeted by G1-arresting rapamycin-receptor complex. Nature 369: 756-758, 1994. PMID: 8008069.
- Brown WS, McDonald PC, Nemirovsky O, Awrey S, Chafe SC, Schaeffer DF, Li J, Renouf DJ, Stanger BZ, and Dedhar S. Overcoming Adaptive Resistance to KRAS and MEK Inhibitors by Co-targeting mTORC1/2 Complexes in Pancreatic Cancer. Cell Rep Med 1: 100131, 2020. PMID: 33294856.
- Buel GR, and Blenis J. CELL SIGNALING. Seeing mTORC1 specificity. Science 351: 25-26, 2016. PMID: 26721988.
- Condon KJ, and Sabatini DM. Nutrient regulation of mTORC1 at a glance. J Cell Sci 132: 2019. PMID: 31722960.
- Crozier SJ, D'Alecy LG, Ernst SA, Ginsburg LE, and Williams JA. Molecular mechanisms of pancreatic dysfunction induced by protein malnutrition. Gastroenterology 137: 1093-1101, 1101 e1091-1093, 2009. PMID: 19427311.
- Crozier SJ, Sans MD, Guo L, D'Alecy LG, and Williams JA. Activation of the mTOR signalling pathway is required for pancreatic growth in protease-inhibitor-fed mice. J Physiol 573: 775-786, 2006. PMID: 16613881.
- Crozier SJ, Sans MD, Wang JY, Lentz SI, Ernst SA, and Williams JA. CCK-independent mTORC1 activation during dietary protein-induced exocrine pancreas growth. Am J Physiol Gastrointest Liver Physiol 299: G1154-1163, 2010. PMID: 20798356.
- Deng L, Chen L, Zhao L, Xu Y, Peng X, Wang X, Ding L, Jin J, Teng H, Wang Y, Pan W, Yu F, Liao L, Li L, Ge X, and Wang P. Ubiquitination of Rheb governs growth factor-induced mTORC1 activation. Cell Res 29: 136-150, 2019. PMID: 30514904.
- Dibble CC, and Manning BD. Signal integration by mTORC1 coordinates nutrient input with biosynthetic output. Nat Cell Biol 15: 555-564, 2013. PMID: 23728461.
- Ding L, Han L, Li Y, Zhao J, He P, and Zhang W. Neurogenin 3-directed cre deletion of Tsc1 gene causes pancreatic acinar carcinoma. Neoplasia 16: 909-917, 2014. PMID: 25425965.
- Elghazi L, Blandino-Rosano M, Alejandro E, Cras-Meneur C, and Bernal-Mizrachi E. Role of nutrients and mTOR signaling in the regulation of pancreatic progenitors development. Mol Metab 6: 560-573, 2017. PMID: 28580286.
- Fingar DC, and Blenis J. Target of rapamycin (TOR): an integrator of nutrient and growth factor signals and coordinator of cell growth and cell cycle progression. Oncogene 23: 3151-3171, 2004. PMID: 15094765.
- Fu W, and Hall MN. Regulation of mTORC2 Signaling. Genes (Basel) 11: 2020. PMID: 32899613.
- Gazzaneo MC, Orellana RA, Suryawan A, Tuckow AP, Kimball SR, Wilson FA, Nguyen HV, Torrazza RM, Fiorotto ML, and Davis TA. Differential regulation of protein synthesis and mTOR signaling in skeletal muscle and visceral tissues of neonatal pigs after a meal. Pediatr Res 70: 253-260, 2011. PMID: 21654549.
- Goberdhan DC, Wilson C, and Harris AL. Amino Acid Sensing by mTORC1: Intracellular Transporters Mark the Spot. Cell Metab 23: 580-589, 2016. PMID: 27076075.
- Hara K, Maruki Y, Long X, Yoshino K, Oshiro N, Hidayat S, Tokunaga C, Avruch J, and Yonezawa K. Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110: 177-189, 2002. PMID: 12150926.
- Hashi M, Yoshizawa F, Onozuka E, Ogata M, and Hara H. Adaptive changes in translation initiation activities for rat pancreatic protein synthesis with feeding of a high-protein diet. J Nutr Biochem 16: 507-512, 2005. PMID: 16043033.
- Hashimoto N, and Hara H. Dietary amino acids promote pancreatic protease synthesis at the translation stage in rats. J Nutr 133: 3052-3057, 2003. PMID: 14519783.
- Javle MM, Shroff RT, Xiong H, Varadhachary GA, Fogelman D, Reddy SA, Davis D, Zhang Y, Wolff RA, and Abbruzzese JL. Inhibition of the mammalian target of rapamycin (mTOR) in advanced pancreatic cancer: results of two phase II studies. BMC Cancer 10: 368, 2010. PMID: 20630061.
- Jewell JL, Russell RC, and Guan KL. Amino acid signalling upstream of mTOR. Nat Rev Mol Cell Biol 14: 133-139, 2013. PMID: 23361334.
- Kim J, and Guan KL. Amino acid signaling in TOR activation. Annu Rev Biochem 80: 1001-1032, 2011. PMID: 21548787.
- Kim J, and Guan KL. mTOR as a central hub of nutrient signalling and cell growth. Nat Cell Biol 21: 63-71, 2019. PMID: 30602761.
- Kong B, Cheng T, Qian C, Wu W, Steiger K, Cao J, Schlitter AM, Regel I, Raulefs S, Friess H, Erkan M, Esposito I, Kleeff J, and Michalski CW. Pancreas-specific activation of mTOR and loss of p53 induce tumors reminiscent of acinar cell carcinoma. Mol Cancer 14: 212, 2015. PMID: 26683340.
- Kong B, Wu W, Cheng T, Schlitter AM, Qian C, Bruns P, Jian Z, Jager C, Regel I, Raulefs S, Behler N, Irmler M, Beckers J, Friess H, Erkan M, Siveke JT, Tannapfel A, Hahn SA, Theis FJ, Esposito I, Kleeff J, and Michalski CW. A subset of metastatic pancreatic ductal adenocarcinomas depends quantitatively on oncogenic Kras/Mek/Erk-induced hyperactive mTOR signalling. Gut 65: 647-657, 2016. PMID: 25601637.
- Laplante M, and Sabatini DM. mTOR signaling in growth control and disease. Cell 149: 274-293, 2012. PMID: 22500797.
- Li W, Zhang H, Nie A, Ni Q, Li F, Ning G, Li X, Gu Y, and Wang Q. mTORC1 pathway mediates beta cell compensatory proliferation in 60 % partial-pancreatectomy mice. Endocrine 53: 117-128, 2016. PMID: 26818915.
- Morran DC, Wu J, Jamieson NB, Mrowinska A, Kalna G, Karim SA, Au AY, Scarlett CJ, Chang DK, Pajak MZ, Australian Pancreatic Cancer Genome I, Oien KA, McKay CJ, Carter CR, Gillen G, Champion S, Pimlott SL, Anderson KI, Evans TR, Grimmond SM, Biankin AV, Sansom OJ, and Morton JP. Targeting mTOR dependency in pancreatic cancer. Gut 63: 1481-1489, 2014. PMID: 24717934.
- Noda T. Regulation of Autophagy through TORC1 and mTORC1. Biomolecules 7: 2017. PMID: 28686223.
- Patel R, Atherton P, Wackerhage H, and Singh J. Signaling proteins associated with diabetic-induced exocrine pancreatic insufficiency in rats. Ann N Y Acad Sci 1084: 490-502, 2006. PMID: 17151324.
- Proud CG. Control of the translational machinery by amino acids. Am J Clin Nutr 99: 231S-236S, 2014. PMID: 24284441.
- Rousseau A, and Bertolotti A. An evolutionarily conserved pathway controls proteasome homeostasis. Nature 536: 184-189, 2016. PMID: 27462806.
- Sabatini DM, Erdjument-Bromage H, Lui M, Tempst P, and Snyder SH. RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs. Cell 78: 35-43, 1994. PMID: 7518356.
- Sancak Y, Bar-Peled L, Zoncu R, Markhard AL, Nada S, and Sabatini DM. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141: 290-303, 2010. PMID: 20381137.
- Sans M, Amin R, Vogel N, D'Alecy L, Kahn R, and Williams J. Specific deletion of insulin receptors on pancreatic acinar cells defines the insulin-acinar axis: implications for pancreatic insufficiency in diabetes. Gastroenterology 140: S-156, 2011.
- Sans MD, Crozier SJ, Vogel NL, D'Alecy LG, and Williams JA. Dietary Protein and Amino Acid Deficiency Inhibit Pancreatic Digestive Enzyme mRNA Translation by Multiple Mechanisms. Cell Mol Gastroenterol Hepatol 11: 99-115, 2021. PMID: 32735995.
- Sans MD, DiMagno MJ, D'Alecy LG, and Williams JA. Caerulein-induced acute pancreatitis inhibits protein synthesis through effects on eIF2B and eIF4F. Am J Physiol Gastrointest Liver Physiol 285: G517-528, 2003. PMID: 12773302.
- Sans MD, Lee SH, D'Alecy LG, and Williams JA. Feeding activates protein synthesis in mouse pancreas at the translational level without increase in mRNA. Am J Physiol Gastrointest Liver Physiol 287: G667-675, 2004. PMID: 15117679.
- Sans MD, Tashiro M, Vogel NL, Kimball SR, D'Alecy LG, and Williams JA. Leucine activates pancreatic translational machinery in rats and mice through mTOR independently of CCK and insulin. J Nutr 136: 1792-1799, 2006. PMID: 16772439.
- Sans MD, Xie Q, and Williams JA. Regulation of translation elongation and phosphorylation of eEF2 in rat pancreatic acini. Biochem Biophys Res Commun 319: 144-151, 2004. PMID: 15158453.
- Saxton RA, and Sabatini DM. mTOR Signaling in Growth, Metabolism, and Disease. Cell 169: 361-371, 2017. PMID: 28388417.
- Schmelzle T, and Hall MN. TOR, a central controller of cell growth. Cell 103: 253-262, 2000. PMID: 11057898.
- Sengupta S, Peterson TR, and Sabatini DM. Regulation of the mTOR complex 1 pathway by nutrients, growth factors, and stress. Mol Cell 40: 310-322, 2010. PMID: 20965424.
- Sinagoga KL, Stone WJ, Schiesser JV, Schweitzer JI, Sampson L, Zheng Y, and Wells JM. Distinct roles for the mTOR pathway in postnatal morphogenesis, maturation and function of pancreatic islets. Development 144: 2402-2414, 2017. PMID: 28576773.
- Su KH, and Dai C. mTORC1 senses stresses: Coupling stress to proteostasis. Bioessays 39: 2017. PMID: 28295473.
- Sung CK, and Williams JA. Cholecystokinin stimulates a specific ribosomal S6 kinase in rat pancreatic acini. Pancreas 5: 668-676, 1990. PMID: 2281080.
- Sung CK, and Williams JA. Insulin and ribosomal protein S6 kinase in rat pancreatic acini. Diabetes 38: 544-549, 1989. PMID: 2653925.
- Torrazza RM, Suryawan A, Gazzaneo MC, Orellana RA, Frank JW, Nguyen HV, Fiorotto ML, El-Kadi S, and Davis TA. Leucine supplementation of a low-protein meal increases skeletal muscle and visceral tissue protein synthesis in neonatal pigs by stimulating mTOR-dependent translation initiation. J Nutr 140: 2145-2152, 2010. PMID: 20962152.
- Vander Haar E, Lee SI, Bandhakavi S, Griffin TJ, and Kim DH. Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat Cell Biol 9: 316-323, 2007. PMID: 17277771.
- von Manteuffel SR, Dennis PB, Pullen N, Gingras AC, Sonenberg N, and Thomas G. The insulin-induced signalling pathway leading to S6 and initiation factor 4E binding protein 1 phosphorylation bifurcates at a rapamycin-sensitive point immediately upstream of p70s6k. Mol Cell Biol 17: 5426-5436, 1997. PMID: 9271419.
- Yang H, Jiang X, Li B, Yang HJ, Miller M, Yang A, Dhar A, and Pavletich NP. Mechanisms of mTORC1 activation by RHEB and inhibition by PRAS40. Nature 552: 368-373, 2017. PMID: 29236692.
- Yang J, Waldron RT, Su HY, Moro A, Chang HH, Eibl G, Ferreri K, Kandeel FR, Lugea A, Li L, and Pandol SJ. Insulin promotes proliferation and fibrosing responses in activated pancreatic stellate cells. Am J Physiol Gastrointest Liver Physiol 311: G675-G687, 2016. PMID: 27609771.
- Yuan HX, Xiong Y, and Guan KL. Nutrient sensing, metabolism, and cell growth control. Mol Cell 49: 379-387, 2013. PMID: 23395268.
- Zhao Y, Schoeps B, Yao D, Zhang Z, Schuck K, Tissen V, Jager C, Schlitter AM, van der Kammen R, Ludwig C, D'Haese JG, Raulefs S, Maeritz N, Shen S, Zou X, Kruger A, Kleeff J, Michalski CW, Friess H, Innocenti M, and Kong B. mTORC1 and mTORC2 Converge on the Arp2/3 Complex to Promote Kras(G12D)-Induced Acinar-to-Ductal Metaplasia and Early Pancreatic Carcinogenesis. Gastroenterology 160: 1755-1770 e1717, 2021. PMID: 33388318.
- Zoncu R, Efeyan A, and Sabatini DM. mTOR: from growth signal integration to cancer, diabetes and ageing. Nat Rev Mol Cell Biol 12: 21-35, 2011. PMID: 21157483.